We use the cotton genus Gossypium as a model system to investigate the mechanistic underpinning of phenotypic evolution and crop domestication in plants.
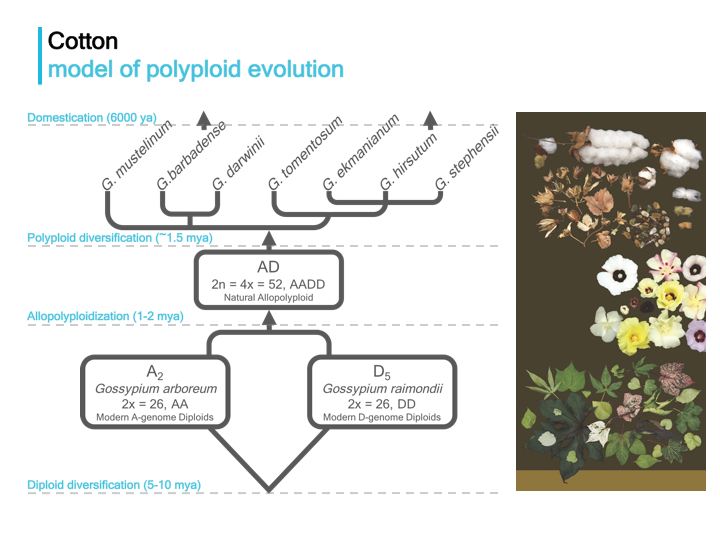
Projects
Unraveling cis-trans interactions and gene regulatory networks underlying salt tolerance in Gossypium
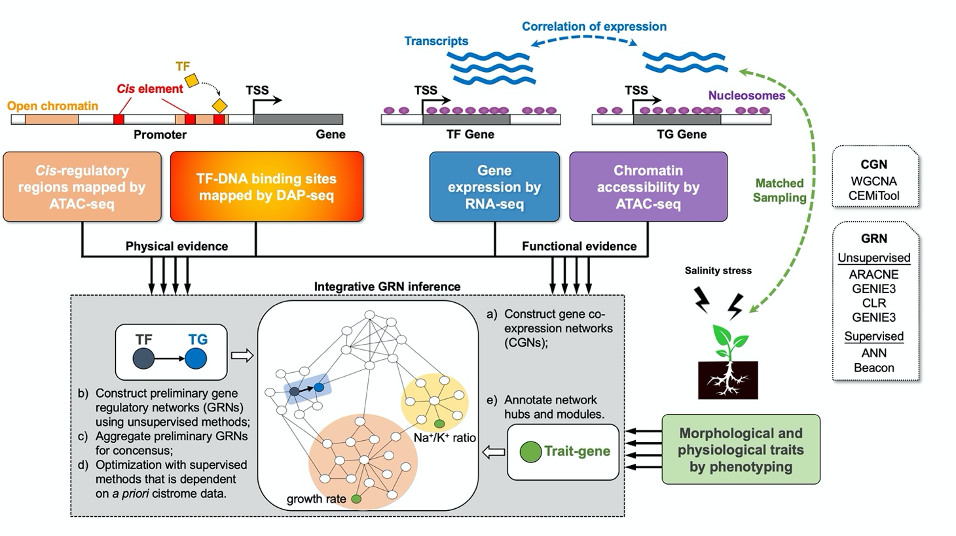
Attempts to develop salinity-tolerant cotton varieties have met with limited success due to the complexity of salt tolerance physiology, high variation in stress response, and especially because of the lack of genetic diversity for tolerance in the narrow gene pool encompassed by elite cultivated lines. Because wild relatives of cultivated cotton have far greater genetic diversity and often naturally occur in high salinity environments, they likely represent an important source of salinity-related germplasm and hold keys to developing a deeper understanding of the relevant biology. This project aims to provide: (1) an enhanced understanding of the phenotypic differences in salt stress adaption among cultivated and wild cotton relatives, thereby identifying elite germplasm and key traits that should be targeted in genetic studies and future breeding programs; (2) detailed and comparative insight into global gene expression and chromatin accessibility dynamics in salt-tolerant wild species relative to salt-sensitive G. hirsutum, thereby providing a detailed view of regulatory divergence and responsiveness related to salt stress; (3) a deep understanding of the cis and trans interactions involved in the regulatory circuits of stress-responsive regulatory gene networks, leading to the discovery of key regulators, thereby facilitating insight into stress biology and facilitating cotton breeding for improved salt tolerance.
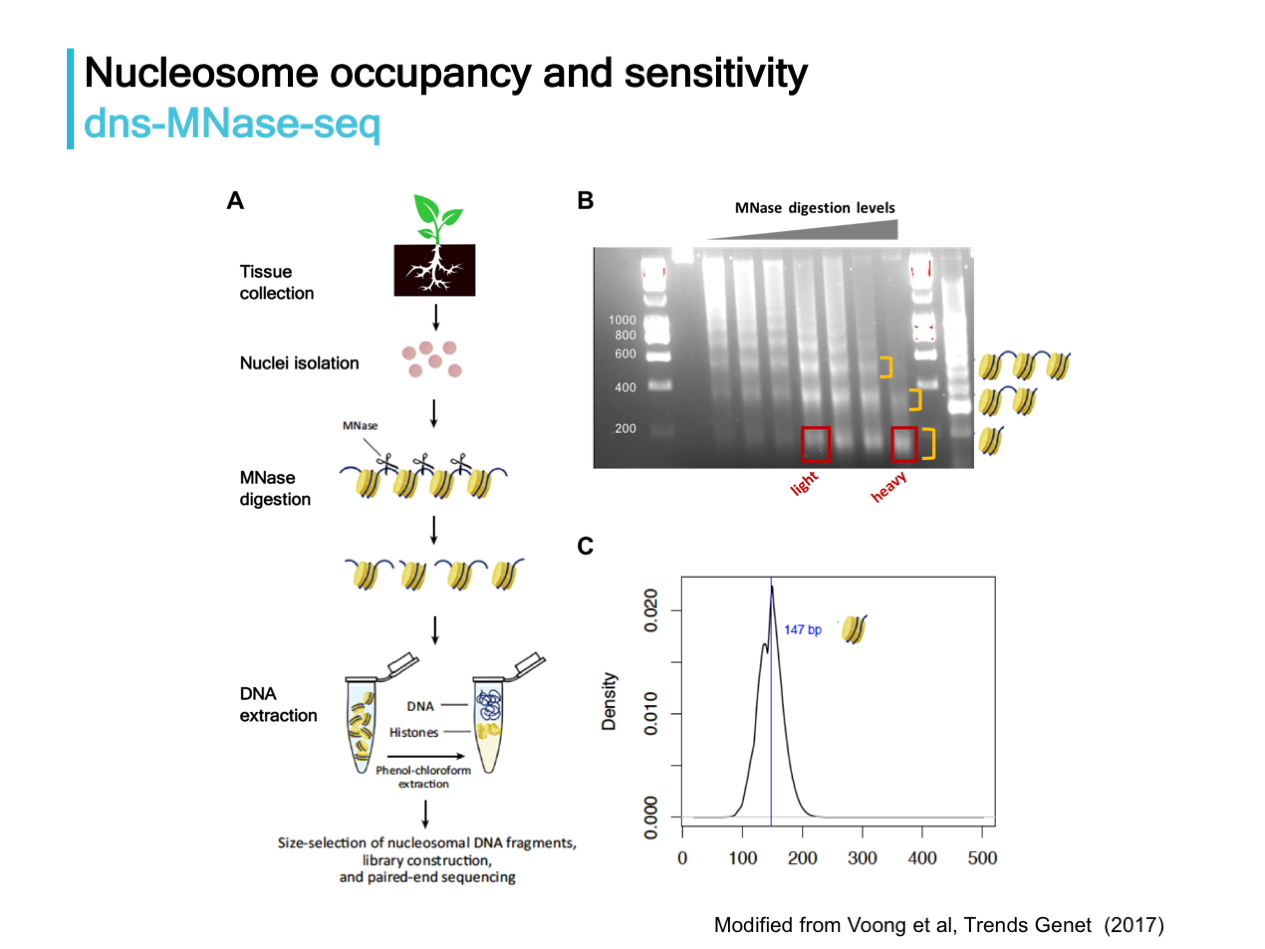
Nucleosome dynamics and duplicated gene expression evolution
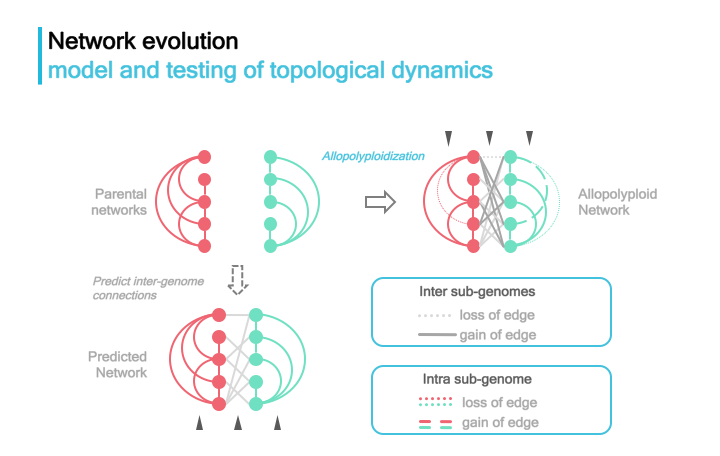
Dynamics of duplicated networks in polyploids
Archived Projects
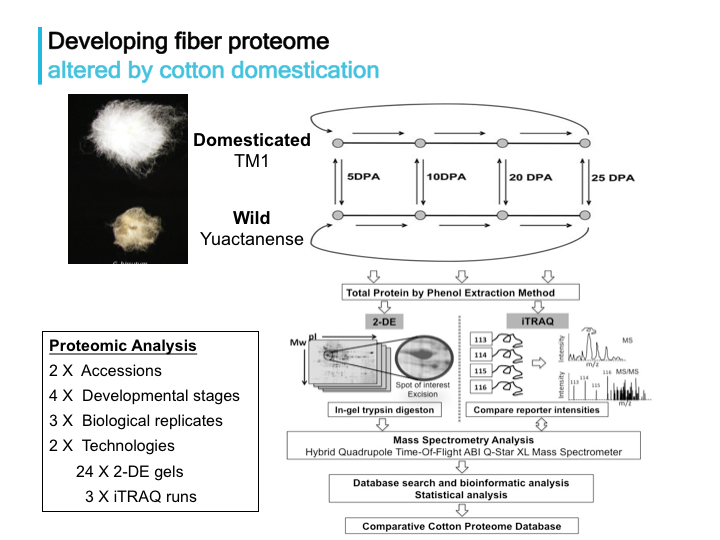
Evolutionary proteomics of cotton: this project was initiated as my phD dissertation work, and has continued to focus on the integration of transcriptome and proteome data.
External Links:
Wendel Lab Reseach |
CottonGen |
Link 3